Borrowed from Chakma J. Genomic Variation: The Search for our Past and Our Future. Journal of Young Investigators. Vol 16, 2009.
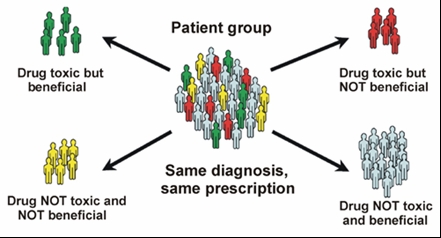
The concept of personalized medicine has received a tremendous amount of media attention as of late, and in this post I thought I would try to outline the various technologies, concepts, and ongoing clinical trials that are coming as a result of these initiatives. To be clear, all medicine is tailored (or should be anyway) to the unique scenario for each patient- but now, the concept of personalized/precision medicine is a bit more specific to treatments. In a nutshell, the idea of “precision” or “personalized” medicine is to try to identify treatments that are likely to be the most helpful to a specific individual IN ADVANCE. By doing this, we could avoid wasting precious time on ineffective therapies and prevent unnecessary side effects. The ability to do this is particularly important in cancer, where some amount of damage to normal and healthy cells is expected from chemotherapy and other anti-cancer treatments.
As we’ve previously discussed, all cancers are not created equal. Within various cancer types, we’ve learned that the genetics and proteins that make up those cancers are incredibly different, affecting the behavior of the tumors, the ability to spread, and response to treatment. Our ability to profile and understand tumor behavior is more advanced in some cancers than others. For example, our understanding has rapidly evolved in breast and lung cancers, whereas in sarcomas, most types still are not well understood with a few exceptions.
For example – breast cancers often are influenced by hormones, including estrogen and progesterone. Additionally, the presence of a protein called Her2neu can be expressed on the surface of breast cancer cells and receive signals that drive abnormal growth. All new breast cancers are profiled by staining for these three proteins – and treatment is adapted to fit the situation. A breast cancer that is estrogen receptor/progesterone receptor positive but Her2 negative may be managed with hormone blockade- using the first targeted therapies, tamoxifen or aromatase inhibitors to block the effect from estrogen. The ability to recognize hormone-receptor positive breast cancers have saved many women with these cancers the need for chemotherapy, instead treating with years of tamoxifen or aromatase inhibitors after their initial surgeries and radiation. Her2 used to be a sign of extremely aggressive breast cancers, but now patients are treated with the anti-Her2 drug Herceptin – and often do well. Breast cancers lacking all three receptors are more aggressive and warrant more intensive chemotherapy. Thus, based on the individual tumor profile, treatment can be adapted for the specific patient, and save some women unnecessary chemotherapy.
Another example is lung cancer. Today, lung cancers can been classified according to mistakes, or mutations in the genetic code, or DNA. Although the pie chart is growing bigger all the time, here is an example from a recent paper showing “actionable” mutations, many of which now have specific drugs that can be quite effective. Patients with EGFR mutations in lung cancer now are often treated first with an oral chemotherapy pill called Tarceva, as opposed to aggressive chemotherapy.
From Pao et al, Nature Medicine, 2012.
The interesting thing is that many of these same mutations show up in other cancers too. BRAF mutations are found in colon cancer and melanoma. But surprisingly, BRAF inhibitors don’t work in colon cancer like they do in melanoma. Cancer is an unstable environment, and new mutations happen all the time, particularly under the pressure of treatments. So something we struggle with is understanding whether a mutation in a cancer is a “driver” mutation, with critical roles in pushing the cancer along the cycle of growth, or a “passenger” mutation, one that happens to be broken but doesn’t really have much significance that we can identify. Using a drug to inhibit a driver mutation is much more likely to be effective than inhibiting a passenger mutation… but it’s tough to know whether it’s a driver or passenger ahead of time.
Thus, there are several companies that offer “sequencing” analysis, which maps the patient’s cancer and looks for mutations in cancer genes. Foundation Medicine and Caris are the two companies I have used to try to look for mutations with potential drugs that might be helpful in a patient’s particular cancer. Additionally, many major cancer centers are now doing their own genome sequencing analyses for patients as part of the initial intake. Conveniently, these tests can be done on tumor samples that are preserved in pathology labs and don’t require new biopsies if the previous sample is large enough.
There are few issues to consider with this testing, however. First, any detected mutations may or may not be significant, given the driver vs. passenger phenomenon we discussed before. Additionally, in the case of a common cancer like colon or lung cancer, there are likely more established treatments with a higher chance of benefit than using a drug specific for a rarer mutation – we simply don’t have any data to support treatment in that scenario. Results need to be interpreted with caution, especially if the test was done on a piece of tumor prior to additional therapies, or in the setting of recurrent or metastatic disease, as these new cancers may have altered their DNA since the initial diagnosis. Finally, the testing is expensive, with Foundation charging over $5500 for their test. Insurance companies are covering this testing more and more, but often they balk at covering the costs for a mutation-specific drug unless there are already clinical trials supporting its use in that particular tumor type. In summary, I use these tests for cancers that are not easily classified (ie, unknown origination sites, or those that don’t neatly fit into either a sarcoma or a carcinoma), cancers without approved and standard treatment options, or in patients that have progressed on standard treatment regimens. I use them to try to guide the patient to a clinical trial whenever possible. Fortunately, Foundation Medicine and Caris reports include lists of potential clinical trials open nationally that might be suitable for those particular mutations detected.
Enter stage right: the upcoming NCI-MATCH Trial and ASCO TAPUR trial. Like the much less-publicized Novartis Signature trial, these studies are HUGE efforts to match patients who have progressed on standard therapies to a large clinical trial providing mutation analysis and then specific drugs for their own mutations. At participating institutions, a patient’s tumor will be sent to NCI for mutation analysis, and then enrolled in a national trial to give them treatment with mutation-specific drugs. They are starting with ten mutation/drug pairs, but expected to expand even more as the trial kicks off. The benefits to these approaches are that the cost of the sequencing analysis will be paid for as part of the study costs, and there are no battles with insurance companies to cover off label uses of drugs.
What about standard chemotherapy? Caris also includes suggestions about chemotherapies likely to be effective or not, based loosely on the genetic profile. Other companies like ChemoFX have also tried to test tumor cells for sensitivity in the lab. Unfortunately, for reasons I’ll get into, cancer cell death in tissue culture (test tubes and Petri dishes) does not always equate to success in human beings. The idea behind these tests is that a biopsy of the tumor is taken and put immediately into cell media in the lab, often times with growth factors and additives, to try to get the cancer cells to grow on plastic. If they take and begin to divide, various drugs are then tested against the new cells. The problems abound… not all of the cells in a tumor will be able to grow on plastic, leading to selection of only a few cells that have the ability to grow – not representing the whole complete tumor in the person. Additionally, as soon as they are exposed to any nutrition or growth factors in the media, the cancer cells adapt their DNA and genes in response to the environment – again altering them from the primary tumor. Finally, when testing drugs in plastic, the usual way that the experiments are conducted is by adding a concentration of drug into the well where the cells are. If you add enough of ANY drug, the cells will die in that scenario. Additionally, there is no barrier between the cells and the drug in plastic – in a human tumor, there might not be enough blood flow to get the drugs inside the whole tumor, or the body’s metabolism may alter or clear the drug long before it arrives at the tumor. So… it’s complicated. Based on all these problems, insurance companies have almost universally refused to cover the costs for these analyses – and no well-controlled studies have showed that adapting treatment based on these results leads to improved survival.
Because of this, others have tried to develop ways to more accurately mimic the entire human tumor for drug testing. Champions Oncology takes tumor samples and implants them in mice, passaging the ones that take into little mouse armies for mini clinical trials of various chemotherapy drugs. This process (called xenografting or PDX) gets around the problem in cell lines, by passaging a larger piece of the tumor. However, while treating mice approximates humans to a better degree than cell lines, there are still issues with getting drug into the tumors. Additionally, most clinical trials in humans are based on stellar data from cell lines and mouse models in the laboratory. Despite working in mice, most drugs still fail once they advance to human trials.
It takes months to generate xenografts, and not all tumors will grow in a mouse. Additionally, it is EXPENSIVE. It costs thousands of dollars to implant the initial tumors, and then there is a charge for each drug tested. No insurance companies have agreed to support it yet either. While the initial data from the company is promising, in most patients the information about potentially promising drugs comes months later, which is a long time to wait – treatment decisions usually will be made before that time. I’ve yet to order this for a patient, but it is intriguing for research and with improved cost containment may make its way into our future efforts in personalizing medicine.
So now with this foundation, you’re all set to go out and browse the numerous compelling articles that are out there about the promise of personalized medicine, and to read the media critically. Fortunately, more and more clinical trials are including analysis of “biomarkers” – so that patients who respond to drugs will be analyzed later for any protein or gene markers that might predict benefit. In the next 5 to 10 years, I have no doubt that patients will walk into my office with a complete profile and analysis of more effective, safer cancer treatment options.


Excellent article!
LikeLike
Great information, Dr. Wilky! Will pass along to a friend of mine being treated for invasive ductal breast cancer, grade 3, stage 1, triple negative hormone receptors and HER2 positive.
LikeLike